Tracking the 2014 Ebola Outbreak Through Its Genes
Genetic detective work also revealed 395 mutations unique to the virus in West Africa
/https://tf-cmsv2-smithsonianmag-media.s3.amazonaws.com/filer/df/54/df549451-aaa0-4765-9642-68b737d4e31d/17775_lores.jpg)
A healer’s funeral was the most likely source of the Ebola virus now spreading in Sierra Leone, one of the hardest hit nations in an outbreak that could infect up to 20,000 across West Africa, according to estimates from the World Health Organization. Tracking the virus involved rapid—and dangerous—genetic detective work that revealed 395 mutations unique to this outbreak. The results, published today in the journal Science, may help scientists discern what makes the virus so severe and inform public health efforts.
Ebola's genome is made of RNA instead of DNA, and it mutates quickly. “Each infection event is a new opportunity to mutate,” says Pardis Sabeti, a geneticist at the Broad Institute at Harvard University in Boston. Five known strains of Ebola infect humans, and the one responsible for this outbreak was identified early on as EBOV, also known as Zaire ebolavirus.
Since its emergence in Guinea in February 2014, this strain of Ebola has spread across West Africa, killing thousands to date. Until this outbreak, Ebola had been restricted to central Africa, making this event unprecedented in both its sheer numbers and its geographic reach. In addition, the virus has been undergoing tiny but potentially significant genetic mutations as it spreads.
Sabeti’s lab has been working on methods to quickly sequence viral samples of Ebola and another hemorrhagic fever, Lassa. When the first whiff of a potential Ebola outbreak hit West Africa, Sabeti put two of her lab members on a plane to Sierra Leone. She and some colleagues at Tulane University had been working with researchers at the country’s Kenema Government Hospital to study Lassa fever. The hospital had a surveillance network in place to pick up Lassa and Ebola cases, and it identified the first case in Sierra Leone, a pregnant woman who miscarried but later recovered.
Kenema researchers retraced the pregnant woman’s steps to the funeral of a healer who had been crossing the nearby border to treat Ebola patients in Guinea. Because they often involve a cleansing of the body, traditional funerary practices can transmit the virus. Sure enough, 13 other women who attended the same funeral were infected with the virus.
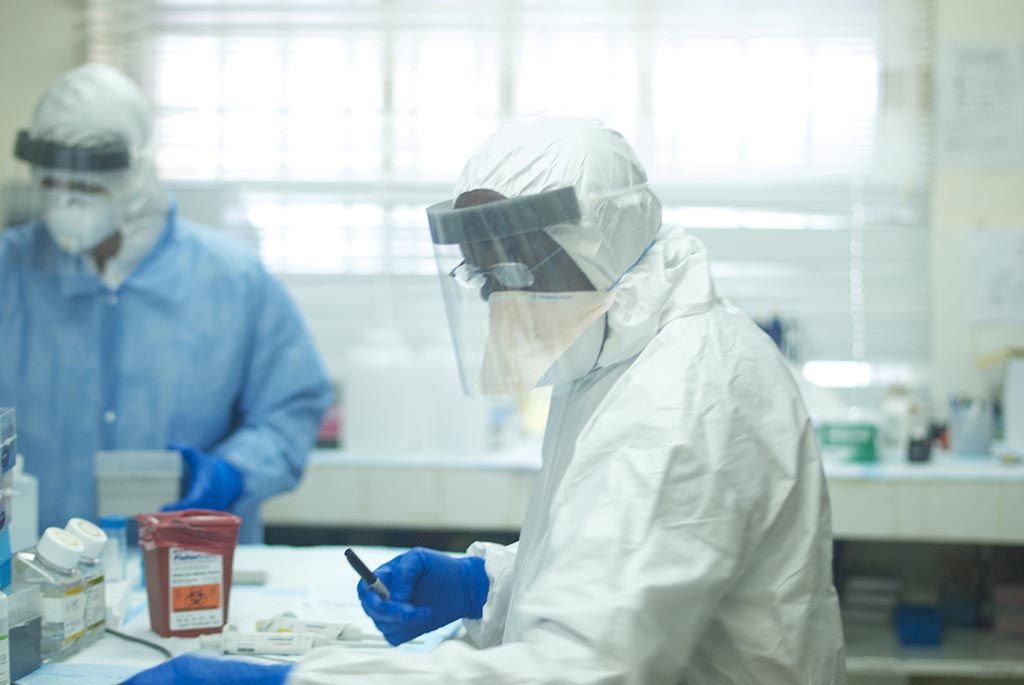
Over the next month, the team collected blood samples from Ebola patients at Kenema and conducted interviews with patients and locals in the field. Unfortunately, this kind of field work comes with serious risk of infection: five of the study’s co-authors contracted Ebola and died prior to publication, including renowned doctor Sheik Humarr Kahn.
Blood samples were then shipped back to Boston, where the virus was rendered inactive and thus safe to handle. Sabeti’s lab then sequenced the samples using a process called deep sequencing, in which they take about 2,000 genetic snapshots of the viral population in the blood.
They came up with 99 viral genomes from 78 patients, and they identified 395 unique mutations. Within Sierra Leone, they could match up differences in the genomes from different patients with data from public health outreach teams at Kenema. “Little mutations will show you the relationships between individual cases,” says Sabeti. Comparing these genomes to those of patients in Guinea and 20 genomes from previous outbreaks, the researchers retraced the virus’s transmission cycles and evolutionary history.
In Sierra Leone, the outbreak stems from the introduction of two genetically distinct viruses, which trace their lineages back to a variant found elsewhere in West Africa around February 2014. From there, it traces its evolutionary history back to the original Ebola outbreak in 1976 in Zaire. Based on its mutation rate, the team thinks the variant could have traveled from central Africa into Guinea as early as 2004, but how it got there is still a mystery. “There are so many missing pieces. It’s kind of like trying to put together an archaeological record,” says Sabeti.
Scientists think that in between outbreaks, Ebola resides in a reservoir of animal populations—likely bats, though the evidence is spotty. Still, the new study backs up evidence that the virus probably only jumped from animals to humans once and has since been transmitted from one human to another. From a public health perspective, that means eating contaminated meat or fruit is less of an issue than human interactions. “You’ve really got to reconsider what you’re telling people, because you don’t want them malnourished when they are most likely to get this from contact [with another human],” says Sabeti.
In any type of outbreak, genetic sequencing can be essential to figuring out exactly what public health workers on the ground are dealing with. So far, that has been logistically very difficult, especially with Ebola. The new work points to the possibility of using genomics, molecular data and on-the-ground interviews together in real time to control the outbreak at hand, notes Roman Biek, who studies infectious disease ecology at the University of Glasgow.
For instance, the findings come on the heels of an announcement this morning that the National Institutes of Health will begin human trials of an Ebola vaccine next week. The nitty gritty genetic details of the cases in Sierra Leone could eventually inform which vaccines, treatments and diagnostic tests would prove most effective in the field.
Diagnostic tests used to confirm cases are based on genetics, and since the team found new mutations in the Sierra Leone cases, it’s possible they might not show up in such lab tests. Sabeti and her colleagues are doing some further digging to say for sure. Some of the mutations flagged in the study could also change the function of viral proteins. Sometimes these mutations produce a working protein, sometimes they don’t. The longer the outbreak goes on, the more functional mutations could pop up, potentially giving the virus an advantage that makes it easier to jump between people.
“That’s one of the biggest fears— that Ebola could become more transmissible than it is now,” says Biek. “It’s important to use genetic data to look at this problem.”
Discerning the exact significance of the mutations will take a massive global effort. Sabeti and her colleagues put all of their raw sequencing data online back in June to make it available to researchers around the world, and other research groups have made similar efforts with sequences from Guinea. Hopefully more teams will follow suit. “We need the entire world to be working on this,” says Sabeti. “I love crowdsourcing, and I think there’s no better use for crowdsourcing than [Ebola].”
/https://tf-cmsv2-smithsonianmag-media.s3.amazonaws.com/accounts/headshot/Screen_Shot_2014-01-27_at_12.05.16_PM.png)






/https://tf-cmsv2-smithsonianmag-media.s3.amazonaws.com/accounts/headshot/Screen_Shot_2014-01-27_at_12.05.16_PM.png)