Cracking the Code of the Human Genome
A Smithsonian magazine special report
Scientists Sequence DNA of Bacteria Responsible for Medieval Leprosy
Genetic information gathered from centuries-old exhumed bones reveals that the infection hasn’t changed much in the past 1,000 years
![]()
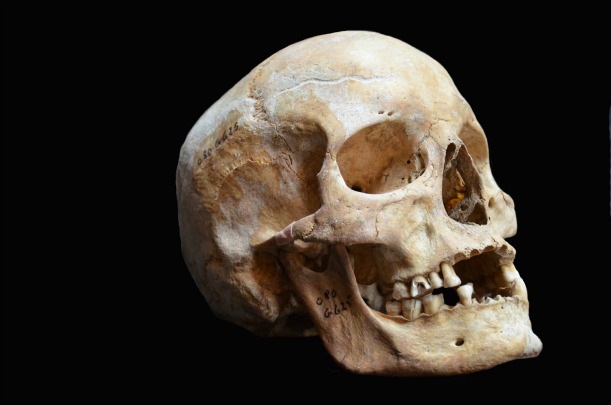
DNA extracted from the skull of this leprosy victim, identified in the study as Jorgen_625, was used to sequence the genome of the medieval. Jorgen_625 lived in Odense, Denmark. Image © Ben Krause-Kyora
For centuries, millions of Europeans suffering from leprosy were shunned by society, made to wear bells that signaled to healthy citizens they were nearby. The infectious illness, also known as Hansen’s Disease, was poorly understood, often believed to be hereditary or a punishment from God. At its height, nearly one in 30 had the disease in some regions; by the 13th century, the number of leper hospitals active in Europe hit its peak at 19,000. Then, in the 16th century, the affliction fell into decline. Soon, it had virtually disappeared from the continent.
The pathogen responsible for leprosy was discovered in 1873 in Norway, squashing previous assumptions about its cause. The earliest written mention of leprosy, one of the oldest-known pathogens to plague humans, appeared in 600 B.C. in China. Historical records show it plagued ancient Greek, Egyptian and Indian civilizations. In 2009, DNA analysis of a first-century man’s remains found in a Jerusalem tomb provided the earliest proven case of leprosy.
Now, DNA sequencing technology has provided clues about the evolution of the bacteria itself. Using well-preserved DNA samples from ancient skeletons, an international team of researchers has sequenced the genome of the pathogen Mycobacterium leprae as it existed in medieval times.
Until now, scientists hadn’t even been able to sequence the pathogen from living people—the bacterium can’t be grown in cell culture in the lab, so scientists usually infect mice with it to achieve a sample big enough for sequencing. The material gleaned from human bones for this study, exhumed from medieval graves, contained a tiny amount of bacterial DNA—less than 0.1 percent, to be in fact. But thanks to extremely sensitive and precise technology, scientists were able to sequence five strains of M. leprae.
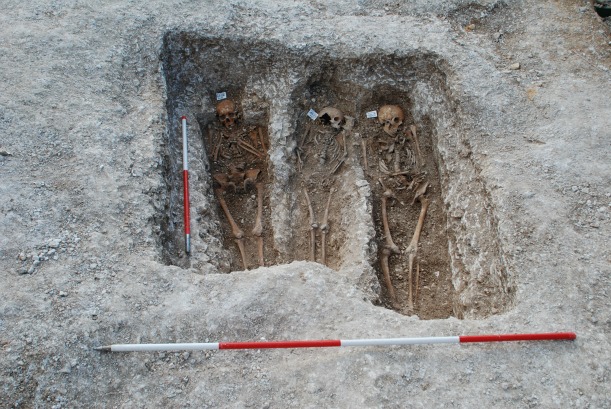

Scientists sequenced DNA found in bones excavated from Medieval graves in Denmark, Sweden and the U.K. Here, remains at the medieval leprosy hospital of St. Mary Magdalen in Winchester, U.K., await excavation. Image courtesy of University of Winchester
Today, more than 225,000 cases of leprosy arise each year, mostly in developing countries. Using samples from some of these cases, the researchers compared the centuries-old sequences to 11 modern strains of the pathogen, extracted from recent biopsies from several geographic regions.
The results, published today in the journal Science, reveal that the bacterium has, in terms of genetic makeup, remained relatively the same despite the last 1,000 years. Only 800 mutations occurred among the 16 genomes in that time, the researchers write. This number means that the mysterious disappearance of the disease by the Middle Ages in Europe can’t be attributed to M. leprae losing its virulence.
“If the explanation of the drop in leprosy cases isn’t in the pathogen, then it must be in the host—that is, in us,” says Stewart Cole, co-director of the study and the head of the École Polytechnique Fédérale de Lausanne’s Global Health Institute. “So that’s where we need to look.”
The pathogen’s genetic resilience was evident in its modern strains. Researchers found that a medieval strain present in Sweden and the U.K. was nearly identical to one currently found in the Middle East. Their findings also suggest that some strains found in the Americas originated in Europe. What they can’t tell us, however, is the direction in which the epidemic spread throughout history.
This research marks a growing trend in using DNA analysis to learn more about epidemics and other devastating events in human history. Last month, scientists sampled 166-year-old Irish potato leaves using similar technology: They determined that a previously unknown strain of P. infestans caused the blight that shrunk 19th-century Ireland’s population by 25 percent. Perhaps future research could someday pinpoint the pathogen responsible for the bubonic plague, commonly known as the Black Death, which wiped out nearly half of Europe’s population between 1347 and 1351.
/https://tf-cmsv2-smithsonianmag-media.s3.amazonaws.com/accounts/headshot/marina-koren-240.jpg)
/https://tf-cmsv2-smithsonianmag-media.s3.amazonaws.com/accounts/headshot/marina-koren-240.jpg)