Scientists Have a New Way of Knowing How Many Sharks Are in the Sea
The predators are elusive, but marine ecologists are finding more of them by analyzing the “environmental DNA” in ocean water samples
/https://tf-cmsv2-smithsonianmag-media.s3.amazonaws.com/filer/18/15/1815e84e-2c8e-4e62-a72e-94c18666c7ae/file-20180502-153878-1p7zmdc.jpg)
Imagine studying animals without seeing them. Does that sound ludicrous? To people like us, who first got interested in biology because we love animals and enjoy studying them, yes, it sounds like a poor deal. Yet, if you think about what forensic investigators do when they seek DNA evidence at a crime scene, or what doctors do when they detect a pathogen in a patient’s blood, it is exactly that: they detect life forms without seeing them.
DNA is life’s blue print. It is present in virtually every organism on Earth, and we usually study it by extracting it from a piece of tissue or a blood sample. But DNA, really, is everywhere: animals shed it constantly, when they scratch themselves, when they release urine, eggs, saliva, excrement and, of course, when they die. Every environment, from your bed to the deepest recesses of the oceans, is full of “biological dust”, mostly cellular material, which contains the DNA of the organisms that left it behind. This, we call “environmental DNA,” or eDNA.
Assisted by increasingly fast, accurate and affordable technology, scientists have begun, in recent years, to sequence this trace DNA from many environments. And this “micro” approach has even proved to be useful to scientists investigating environments as vast as the oceans.

Many marine animals are large, rare, elusive and highly mobile. Sharks are an obvious example: in the oceans they make up a small proportion of the biomass, most of them are pretty difficult to catch, and they have been in conflict with humans since we started venturing at sea. With a few exceptions, they avoid us, and because of us many have become threatened with extinction.
This is why we thought it would be interesting to see if, just by sampling a few bottles of ocean water (and the DNA fragments therein), we could rapidly map shark presence and distribution, without engaging in wild chases or employing time and resource-intensive shark fishing methods. We were happy to find out that, indeed, this was possible, and that different species could be detected in different geographical regions, although the areas that had been more affected by humans would show scant presence of sharks.
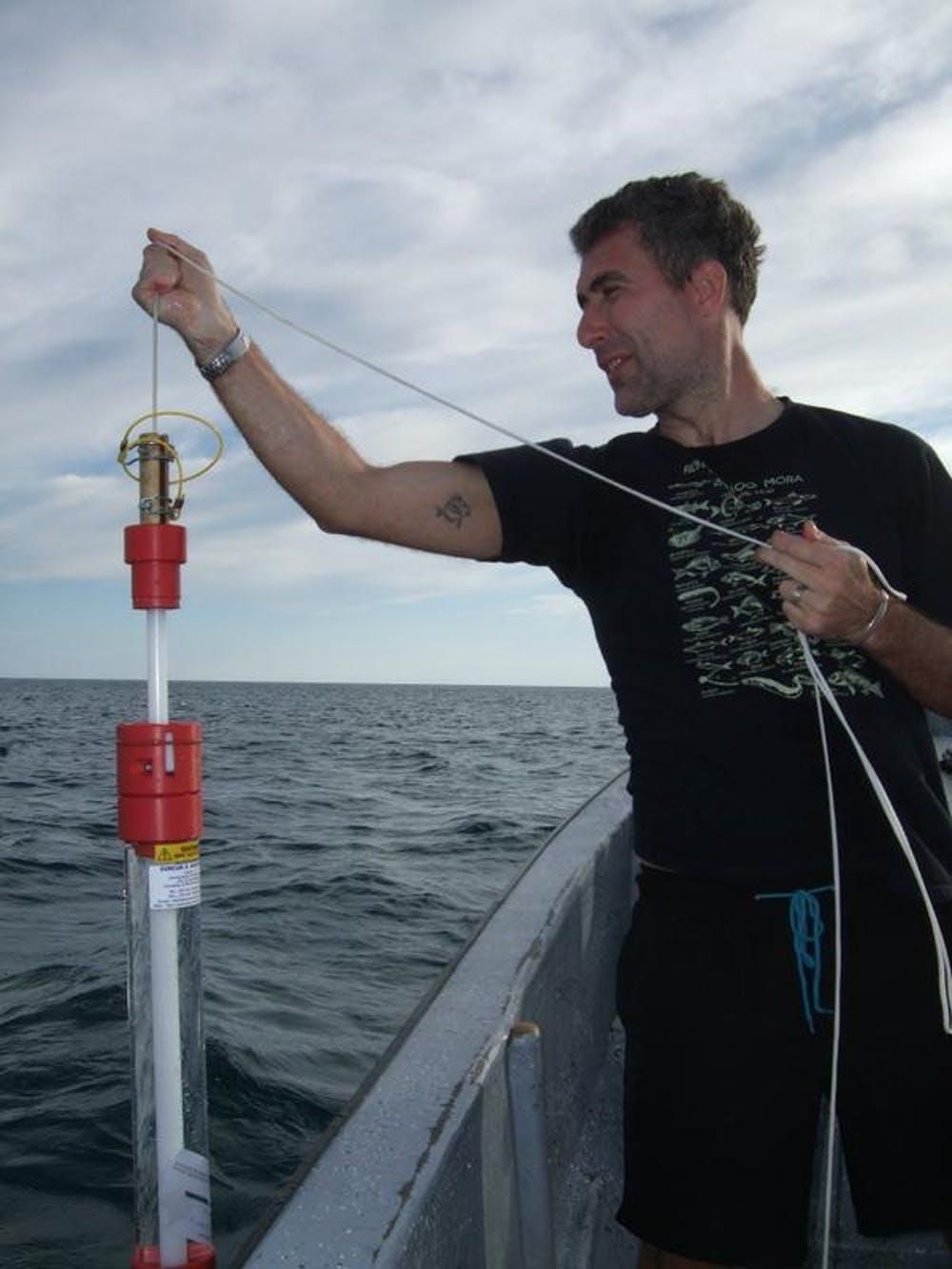

But the true measure of the efficiency of this eDNA approach to shark monitoring would only be revealed when contrasted against established, tried-and-tested methodologies, such as scuba-diving visual censuses or baited underwater camera recordings.
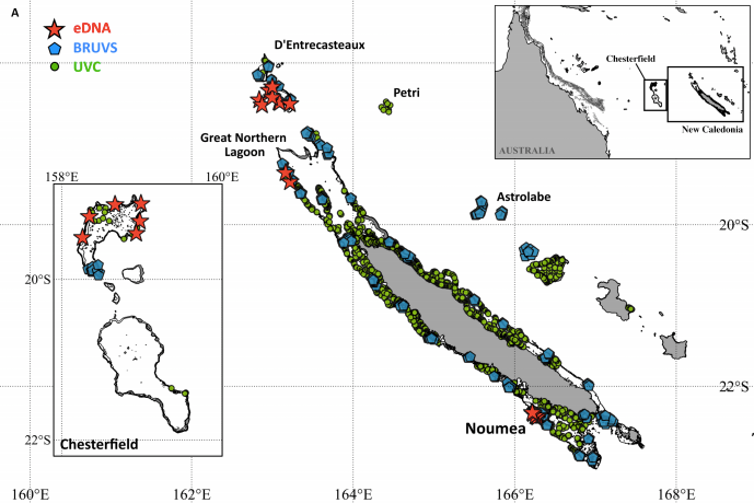
This was the focus of our most recent study, conducted with colleagues based in the South Pacific archipelago of New Caledonia, France, Australia and the U.S., and now published in the journal Science Advances. The results were very exciting: 22 water samples collected over a few weeks detected more sharks than hundreds of baited underwater camera observations over two years, and thousands of scuba dives over a period of decades. Nearly half of the species detected through environmental DNA could not be found at all using traditional methods. And while eDNA could detect the presence of some sharks in about 90 percent of the samples, underwater cameras could only manage just over 50 percent, and scuba diving around 15 percent.
Interestingly, eDNA outperformed the other methods in both pristine and impacted areas. A range of shark species were detected even in busy, noisy and depleted areas, where they were thought to be extirpated. This suggests some “dark diversity” may still be present, in the form of remnant individuals and groups requiring protection. Similarly, eDNA can help by revealing the appearance of newly established, alien species that are expanding their range. All of this is good news for everyone, and this is why.
Given the speed and efficiency of eDNA sampling, a much larger portion of the sea can be screened, in a shorter time, to gather an overview of the patterns of diversity across large areas and habitats, along various environmental gradients, and at different times. Potentially, we could rapidly build maps of species diversity and use them to create predictive models and identify the factors that influence diversity, while methods are being developed to improve the quantitative aspect of eDNA detection, also in other charismatic species. All of it will be of great help to those who must devise plans to protect crucial habitats and ecosystems.
Environmental DNA science is still rapidly developing. The databases that we use to match the unknown sequences retrieved from the sea must be enriched with new DNA references of many existing species – every multi-species eDNA study to date has detected large amounts of sequences that could not be matched against any reference. A significant proportion of these belong to organisms that are yet to be described by scientists.
The “DNA probes” currently available will have to become longer, as short sequences may sometimes fail to distinguish closely related species. For instance, the blacktip shark shared some identical sequences with the grey reef shark along the DNA stretch used in our study. Nevertheless, all the initial indications suggest that this approach can get us a step closer to understanding and better managing the largest ecosystem on Earth.
This article was originally published on The Conversation.
Stefano Mariani, Chair in Conservation Genetics, University of Salford
Judith Bakker, Research Fellow, Environment & Life Sciences, University of Salford