How DNA Testing Can Tell You What Type of Fish You’re Really Eating
By analyzing a the DNA of fish sold across the country, researchers have found that roughly a third of U.S. seafood is mislabeled
/https://tf-cmsv2-smithsonianmag-media.s3.amazonaws.com/filer/20130820094131fish-copy.jpg)
The menu says red snapper, but it’s actually tilapia. The white tuna, meanwhile, is really escolar, while the seabass is Antarctic toothfish.
Welcome to the wild world of modern seafood, where not everything is as it seems. New research is revealing that merchants and fish dealers often mislabel their product as an entirely different species to fetch a better price at market. A study realeased last week by UK researchers found that a number of species in the skate family are sold as “sting ray wings,” while a separate study produced in February by the group Oceana found that, of 1215 seafood samples from 674 restaurants and grocery stores in 21 U.S. states, a full third were mislabeled. In Chicago, New York, and Washington, DC, every single sushi bar that was tested was found to sell at least one mislabeled fish species.
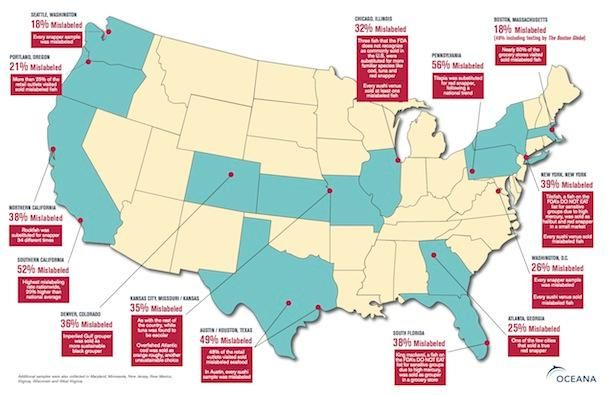
How did the researchers figure all this out? Through the innovative use of DNA barcoding, in which a specific segment of genetic material (analogous to a product’s barcode) in a piece of fish is used to determine exactly which species it truly belongs to. For years, we had no real way of determining the true species of a piece of seafood—a filet of fish, after all, often looks like any other filet—but this new application of an existing scientific technique is rapidly becoming a crucial tool in combating seafood fraud.
Testing a piece of fish to determine its species is fairly straightforward—scientists perfected DNA barcoding years ago, albeit typically as part of other sorts of projects, like cataloging the complete assortment of species in a given ecosystem. Analyzing the DNA in a piece of fish is a relatively similar process.
To start, researchers acquire a piece of fish and freeze it, as fresher and better-preserved tissue samples generally yield more accurate results. Then, in the lab, they slice off a tiny piece of the sample for testing.
To extract and isolate the DNA from the tissue, scientists break open the cells—either physically, by grinding them or shaking them in a test tube filled with tiny beads, or chemically, by exposing them to enzymes that chew through the cell membrane. Next, they remove other components of the cell with various chemicals: proteases digest proteins, while RNAase digests RNA, an alternate form of genetic material that could cause errors in DNA testing if left in place.
Once these and other substances are removed, the remaining sample is put in a centrifuge, which spins it at high speed so that the densest component—in this case, DNA—is isolated at the bottom of the tube in a pellet. A variety of different approaches are currently used to sequence the DNA, but all of them achieve the same end—determining the sequence of base pairs (the building blocks of DNA that are unique to each organism), at one specific location in the fish’s genome. All fish of the same species share the same sequence at that location.
As part of broader DNA barcoding projects, other scientists have analyzed the sequence of base pairs at that same genetic location in thousands of pieces of fish tissue that can definitively linked to species. Thus, by comparing the genetic sequence in the mystery fish tissue to databases of other species’ known genetic sequences, such as FISH-BOL (which stands for Fish-Barcode Of Life and contains the barcodes of 9769 fish species so far), scientists can tell you if, say, the grouper you thought you were buying was actually Asian catfish.
Figuring out which species a piece of fish truly belongs to has significance that goes far beyond gastronomy. For one, cheaper fish species are most often substituted for more expensive ones: tilapia, which goes for around $2.09 per pound, is billed as red snapper, which can commonly fetch $4.49 per pound. (The fact that inexpensive fish is so commonly passed off as a pricier variety, while the reverse occurs much more rarely, indicates that intentional mislabeling by sellers is at play, rather than innocent misidentification.)
Additionally, species that are dangerously overfished and are on the verge of ecological collapse—such as orange roughy—are sometimes substituted for more environmentally-benign varieties. Customers that make the effort to choose sustainable types of seafood, in these cases, are thwarted by mislabeling.
Eating different species can also have vastly different effects on your own health. For one, different fish species can have different fat and calorie contents, so mislabeling can lead the nutrition-conscious astray. Moreover, certain species, like tilefish, are on the FDA’s “do not eat” list for sensitive groups of people (such as pregnant women) because of their high mercury content. The Oceana study, though, found several instances of tilefish being sold as red snapper. Perhaps even worse, 94 percent of the white tuna tested in the study was actually a fish called escolar, which has been found to contain a toxin that when ingested, even in small quantities, can cause severe diarrhea.
So, what to do? Testing the fish’s DNA at home is probably beyond most people’s capabilities. So to avoid being duped, Oceana recommends asking sellers lots of questions about a fish’s origin, scrutinizing the price—if a fish is being sold far below market value, it’s probably mislabeled as a different species—and buying whole fish at markets when possible.
/https://tf-cmsv2-smithsonianmag-media.s3.amazonaws.com/accounts/headshot/joseph-stromberg-240.jpg)
/https://tf-cmsv2-smithsonianmag-media.s3.amazonaws.com/accounts/headshot/joseph-stromberg-240.jpg)